Recommended videos
Topic: VMD: Preparing, Analyzing, and Visualizing Molecular Dynamics Simulations
Presenters: John Stone, University of Illinois Urbana-Champaign and Abhishek Singharoy, Arizona State University, Duration: 1 hour, 23 minutes.
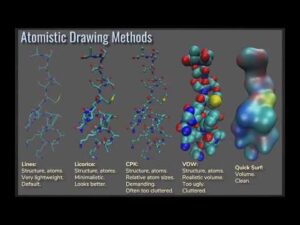

This webinar takes a brief introductory overview of some of the basic features and capabilities of VMD for molecular visualization.
VMD is a molecular visualization program for displaying, animating, and analyzing large biomolecular systems using 3-D graphics and built-in scripting. It provides a wide variety of methods for rendering and coloring a molecule: simple points and lines, CPK spheres and cylinders, licorice bonds, backbone tubes and ribbons, cartoon drawings, and others. VMD can also be used to animate and analyze the trajectory of a molecular dynamics (MD) simulation. In particular, VMD can act as a graphical front end for an external MD program by displaying and animating a molecule undergoing simulation on a remote computer. Duration; 56 minutes.

In this one-hour webinar, we show some VMD (Visual Molecular Dynamics) features that go beyond the basics covered in our first VMD webinar. After a short summary of VMD fundamentals, we learn to:
- Navigate molecular trajectories (as opposed to single structures)
- Produce basic movies using the “VMD Movie Maker”
- Control VMD from text scripts (basic programming)
- Use scripts and external programs together to produce customized movies from molecular trajectories in an automated fashion
