Picture by Sebastian Caro Ortiz
Announcement "Molecular Simulations Workshop: Hands-On Training with RASPA"
iRASPA/RASPA
Recent News
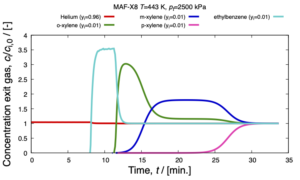
RUPTURA
We announce RUPTURA, soon to be released. It will be a free and open-source software package for (i) the simulation of breakthrough curves, (ii) mixture prediction using methods like the Ideal Adsorption Solution Theory (IAST) and segregated-IAST, and (iii) fitting of isotherm models on computed or measured adsorption isotherm data. Breakthrough plots and movies of the column properties are automatically generated.
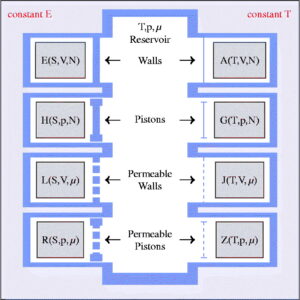
RASPA 3
RASPA3 is in progress. It will be available end of 2024. Redesigned and reimplemented from the ground up in C++23, it will contains many code and speed improvements. Preliminary test shows a five-fold increase in speed for adsorption isotherms of small molecules in rigid nanoporous materials. The input file formats are kept similar to RASPA2, but output files have been significantly improved. On launch, it will be freely available from GitHub.

UvA Researchers Collaborate on New Handbook of Porous Materials
A group of authors including researchers and alumni from the Van ‘t Hoff Institute for Molecular Sciences (HIMS) have just completed a four-volume reference work covering the fundamentals and key applications of porous materials. This “Handbook of Porous Materials” will be published in November by World Scientific Publishers in a digital and a print edition.
Recent Workshops/Schools
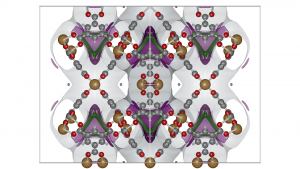
USA RASPA School/Workshop
The USA RASPA workshop is held at University of Massachusetts (Amherst, MA, USA) on 15 June – 18 June 2026 and focuses on a practical understanding of molecular simulations of fluids, ionic liquids, and nanoporous materials and applying the RASPA molecular simulation code to practical examples. The duration is 3.5 days, with lectures in the morning and exercises in the afternoon. Deadline: May 29, 2026

iRASPA/RASPA/RUPTURA Online Workshop 2025
One-day online (via teams) iRASPA/RASPA workshop on Friday September 12, 2025. Times: 9h00 – 17h30 CET (Central European Time). This workshop/school focuses on a practical understanding of visualization and molecular simulation of nanoporous materials and fluids, using (i)RASPA and RUPTURA for breakthrough computations.
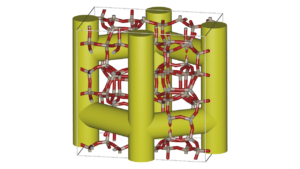
iRASPA/RASPA workshop Delft 2024
Announcement: The (i)RASPA workshop is held at Delft University of Technology (The Netherlands), 2,3,4 September 2024, just before the start of the Thermodynamics2024 conference (thermodynamics2024.org) at Delft. The workshop focuses on a practical understanding of molecular simulations of fluids, ionic liquids, and nanoporous materials and application to practical examples. The duration is 3 days, with morning-lectures and afternoon-exercises.
Recent Blog Posts
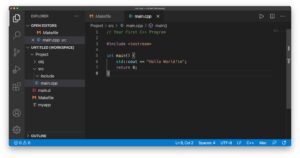
Visual Studio Code C/C++/Fortran with CMake
Visual Studio Code is a free source-code editor made by Microsoft for Windows, Linux and macOS. C/C++ support for Visual Studio Code is provided by a Microsoft C/C++ extension to enable cross-platform C and C++ development on Windows, Linux, and macOS. The Code Runner extension allows execution of single files. For project compilation, consisting of multiple files, the C/C++ Makefile Project extension can be used.
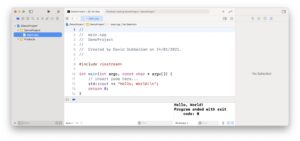
Best C/C++ Integrated Development Environment for Beginners
Settings up your computer for C++ programming can be daunting. For beginners, Xcode is the best IDE for macOS, the Visual Studio IDE is one of the most popular and best IDE on windows, and QtCreator is a good choice on Linux. Installation and setup is straightforward, but the IDEs require significant disk storage.

Visual Studio Code C/C++/Fortran with Multiple Source Files
Visual Studio Code is a free source-code editor made by Microsoft for Windows, Linux and macOS. C/C++ support for Visual Studio Code is provided by a Microsoft C/C++ extension to enable cross-platform C and C++ development on Windows, Linux, and macOS. The Code Runner extension allows execution of single files. For project compilation, consisting of multiple files, the C/C++ Makefile Project extension can be used.
